Display Protein Structure with 3Dmol.js
How to display a protein structure with 3Dmol.js?
✍: FYIcenter.com
![]() You can follow these steps to display a protein structure by loading
an online PDB file.
You can follow these steps to display a protein structure by loading
an online PDB file.
1. Get URL of the online PDB file:
https://files.rcsb.org/download/1YCR.pdb
2. Create an HTML file, View-PDB-Online.html, with the following code. The PDB file loaded with an AJAX call.
<html><head>
<script src="/3Dmol-min.js"></script>
<style>
.mol-container {width: 400px; height: 400px; position: relative;}
</style>
</head>
<body>
<div id="container-01" class="mol-container"></div>
<script>
$(function() {
let element = $('#container-01');
let config = { backgroundColor: 'grey' };
let viewer = $3Dmol.createViewer( element, config );
let pdbUri = 'https://files.rcsb.org/download/1YCR.pdb';
jQuery.ajax( pdbUri, {
success: function(data) {
let v = viewer;
v.addModel( data, "pdb" ); /* load data */
v.setStyle({}, {cartoon: {color: 'spectrum'}}); /* style all atoms */
v.zoomTo(); /* set camera */
v.render(); /* render scene */
},
error: function(hdr, status, err) {
console.error( "Failed to load PDB " + pdbUri + ": " + err );
},
});
});
</script>
</body></html>
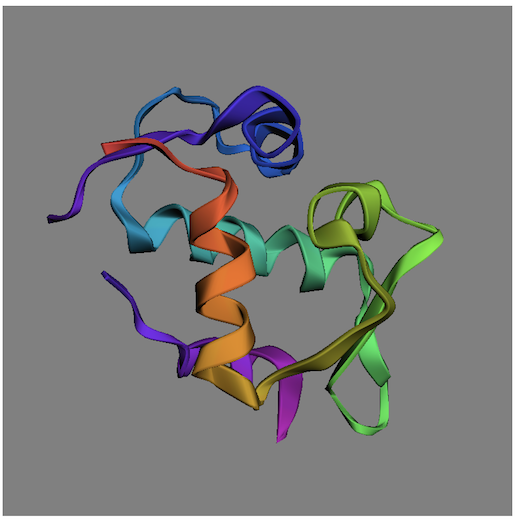
3. View the HTML document through your Web server: http://localhost/Display-Molecule.html. You see the protein structure displayed. You can room in and rotate it with your mouse.
⇒ Use of Anonymous Function for 3Dmol.js
⇐ Display 3-D Molecule Structure with 3Dmol.js
⇑ Getting Started with 3Dmol.js
⇑⇑ 3Dmol.js FAQ
2022-12-26, 1820🔥, 0💬