"pdb=..." - Load PDB Data in 3Dmol Viewer
How to load a protein structure with the PDB ID in the Online 3Dmol Viewer?
✍: FYIcenter.com
![]() You can use the pdb={PDB_ID} URL parameter to load protein data from
https://www.rcsb.org/ in the Online 3Dmol Viewer.
You can use the pdb={PDB_ID} URL parameter to load protein data from
https://www.rcsb.org/ in the Online 3Dmol Viewer.
https://3dmol.org/viewer.html?pdb={PDB_ID}
For example, enter the following URL in Web browser. You see the protein structure is displayed in the 3Dmol Viewer.
https://3dmol.org/viewer.html?pdb=1YCR
By default the structure is displayed in the "line" style as shown below.
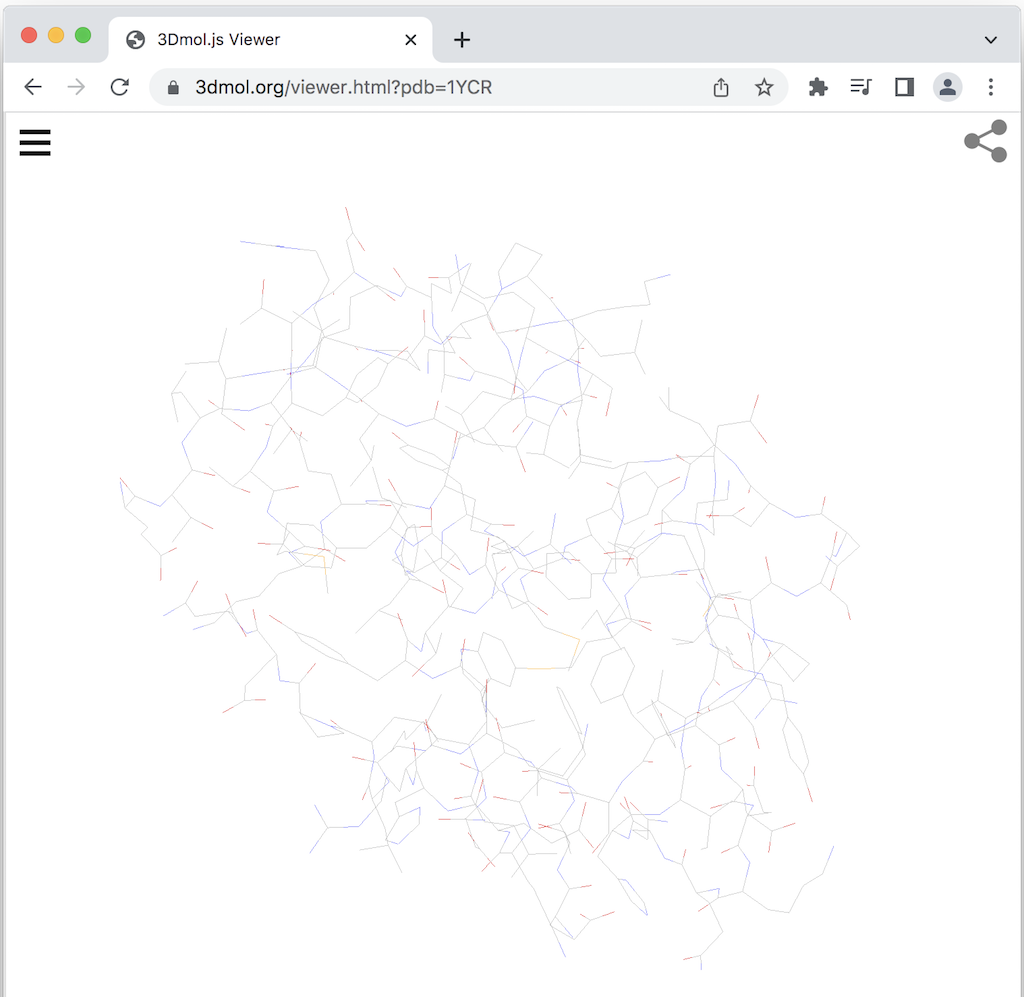
Note that the protein structure data is loaded with the following PDB API:
http://files.rcsb.org/view/{PDB_ID}.pdb
⇒ "style=..." - Specify Display Style in 3Dmol Viewer
⇐ URL Parameters for 3Dmol Viewer
⇑ Using Online Server of 3Dmol Viewer
⇑⇑ 3Dmol.js FAQ
2023-01-06, 618🔥, 0💬